Spatialproteomics: an interoperable toolbox for analyzing highly multiplexed fluorescence images#
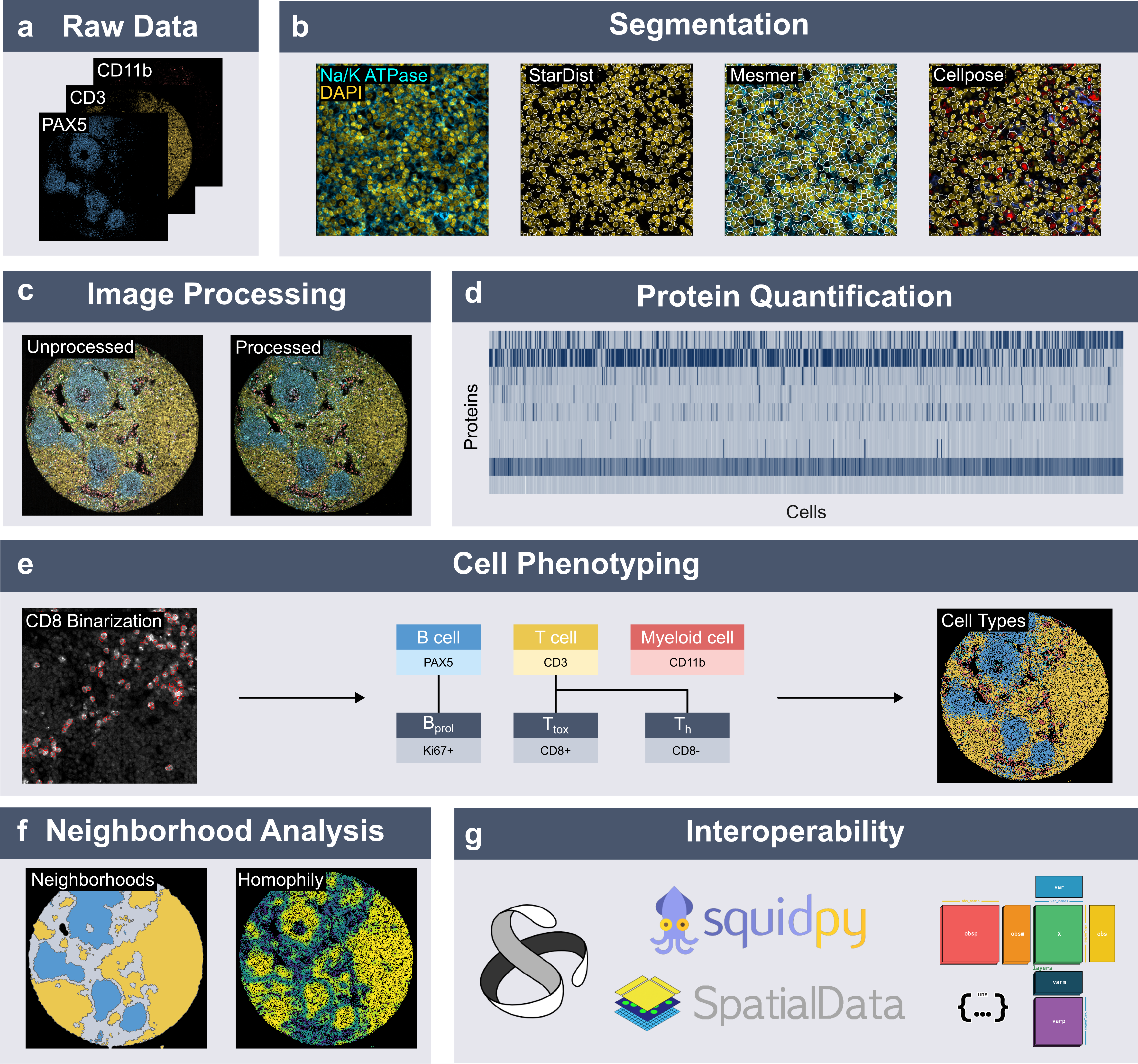
Spatialproteomics is an interoperable toolbox for analyzing high-dimensional imaging datasets. This analysis involves a sequence of steps, including segmentation, image processing, marker quantification, cell type classification, and neighborhood analysis. Built on top of xarray and dask, spatialproteomics can process images that are larger than the working memory. It supports synchronization of shared coordinates across data modalities such as images, segmentation masks, and expression matrices, which facilitates easy and safe subsetting and transformation.
Tutorials:
- Installation
- Example Workflow with spatialproteomics
- Example Workflow with spatialproteomics and spatialdata
- Plotting
- Segmentation
- Image Processing
- Cell Type Prediction
- Neighborhood Analysis
- Interoperability
- Downstream Analysis
- Interactivity with napari
- Customizability
- Subselecting Data
- Interactive Removal of Artifacts
- Integration of Multiple Technologies
- FAQ

